Scientists have discovered new vulnerabilities in a protein that’s often called the “Death Star” of cancer, opening up the possibility of new drugs to target it. This could be a game-changer, as the protein – KRAS – is one of the most frequently mutated cancer proteins, often found in the deadliest cancer types.
The rest of this article is behind a paywall. Please sign in or subscribe to access the full content.KRAS mutations are found in one in 10 human cancers. Although the protein was first discovered over 40 years ago, only two drugs that directly target it have so far been approved for clinical use. This resistance to drug development efforts, as well as its spherical shape, earned it the nickname of the “Death Star” protein.
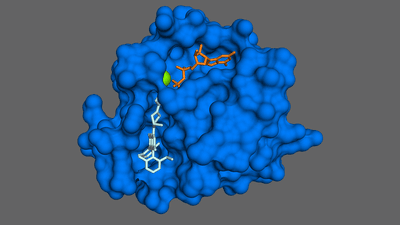
The chief target for drugmakers when they want to deal with a protein like KRAS is its allostery communication system. Think of a lock and key: there are various “locks” secreted all over the surface of a KRAS molecule, waiting for the right “key” in the form of another chemical compound that can bind to the “lock” site.
Once these “keys” bind, the KRAS protein will undergo a change in its shape that can subsequently affect how it behaves. Developing drugs that bind to KRAS at the right sites, therefore, can allow scientists to take control of its activity. These types of drugs can be highly specific to KRAS, minimizing the risk of side-effects, and can allow us to manipulate the protein in very subtle and targeted ways.
That all sounds great, but there’s a big problem: finding the allosteric sites in the first place. The scientific literature contains more than 300 published structures of the KRAS protein, but that’s still only led to the successful development of two drugs, sotorasib and adagrasib. Both of these bind to pockets on the surface of the protein, preventing the binding of other compounds that could activate it. What we really need, then, is more pockets.
Now, a team of researchers at the Centre for Genomic Regulation in Spain and the Wellcome Sanger Institute in the UK have come up with a new method of mapping all the possible allosteric sights at once.
“In this study we demonstrate a new approach that can map allosteric sites systematically for entire proteins. For the purposes of drug discovery, it’s like turning the lights on and laying bare the many ways we can control a protein,” explained co-author André Faure in a statement.
“The unique selling point of our method is its scalability. In this work alone we made more than 22,000 biophysical measurements, a similar number as the total ever made for all proteins before we started harnessing the remarkable strides in DNA sequencing and synthesis methodologies,” added first author Chenchun Weng.
By creating over 26,000 variants of the KRAS protein, manipulating just a couple of its amino acids at a time, the researchers were able to systematically check how each variant responded to six potential binding proteins, including the ones that are critical for causing cancer. With the help of artificial intelligence (AI) software, they revealed that KRAS has many more allosteric sites than we ever suspected.
One that caught the team’s attention is known as “pocket 3”. It hasn’t received much attention in previous pharmaceutical research, but they believe it is worth exploring. This first-ever complete map of allosteric sites in a protein could also generate new ways of targeting only mutated KRAS proteins in cancer, sparing healthy versions of the protein and hopefully decreasing side-effects.
“The big challenge in medicine isn’t knowing which proteins are causing diseases but not knowing how to control them,” said senior author Ben Lehner.
“Our study represents a new strategy to target these proteins and speed up the development of drugs to control their activity. The nature of targeting allosteric sites means that the resulting drugs are likely to be safer, more effective treatments than the ones we have right now.”
The study is published in Nature.