Understanding embryonic development is critical to understanding how congenital defects occur. Scientists have been able to view individual cells during development for quite some time, but they lacked the ability to track cells as they migrate through the body until now. The research was led by Philipp Keller of the Howard Hughes Medical Institute’s Janelia Research Campus and the results were published in Nature Methods.
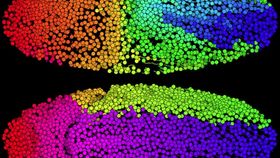
The rest of this article is behind a paywall. Please sign in or subscribe to access the full content.Keller’s team developed a simultaneous multi-view (SiMView) light sheet microscope in 2012 that could obtain 3D images from a subject over the course of several days, which allowed them to simultaneously observe 20,000 cells as they change and migrate during development. This latest research has allowed them to better understand the immense amount of data and target the nervous system specifically, viewing it at the earliest stage possible.
Different species obviously develop at different rates. While a fruit fly embryo develops its nervous system in a day, zebrafish take about two days, and a mouse takes a full week. The SiMView obtains tens of thousands of data points over that time, which makes a tall order out of the data’s analysis. In order to make sense of everything, they developed a computer program in order to track cell movement as the microscope collects the images.
“We want to reconstruct the elemental building plan of animals, tracking each cell from very early development until late stages, so that we know everything that has happened in terms of cell movement and cell division,” Keller explained in a press release.
“In particular, we want to understand how the nervous system forms. Ultimately, we would like to collect the developmental history of every cell in the nervous system and link that information to the cell's final function. For this purpose, we need to be able to follow individual cells on a fairly large scale and over a long period of time.”
The computer is being asked to interpret several terabytes of data in relatively short order, and the system can have difficulty identifying densely-packed cells have different shapes and are moving in different directions. In order to simplify the process, each 3D pixel (called a voxel) was sorted into larger groups of “supervoxels,” which makes the images 1000 times more simple to interpret. The computer can then identify the nucleus of a cell within each supervoxel, allowing the scientists to track individual cells and view their lineage over time.
“You know the path, you know where it is at a certain time point. You know it divided at a certain point, you know the daughter cells, you know what mother cell it came from,” Keller says.
After studying the cell lineage of nearly 300 neuroblasts in fruit flies, the team found that they could predict how movement of cells early in development would affect that cell’s fate. Analyzing cell migration was also successfully used in zebrafish and mice, with a 97% linkage accuracy between the species. Subsequent research will first tackle longstanding questions about embryonic development. Keller’s team hopes others will begin to study specific aspects of development as well, and have made their amazing software open access to facilitate the growth of this field of study.