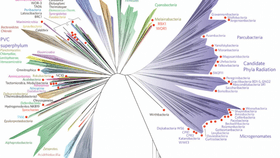
The addition of bacteria that have never been grown in the lab and are only known about from sequencing their DNA has radically expanded the tree of life. These uncultivatable microbes are now thought to account for up to a third of all biodiversity, massively swamping all the species we can see and usually think make up the majority of our ecosystems.
The rest of this article is behind a paywall. Please sign in or subscribe to access the full content.“This is the first three-domain genome-based tree to incorporate these uncultivable organisms, and it reveals the vast scope of as yet little-known lineages,” explains Jill Banfield from UC Berkeley, who co-authored the paper published in Nature Microbiology. It now turns out that they dominate the main domains of life on this planet, which are split into three: Bacteria, Archaea, and Eukaryota. You, me, and most livings things we can see with the naked eye are part of Eukaryota, while the other two domains are formed of single-celled organisms, with subtle differences separating the Bacteria and Archaea.
The new tree reconfirms that the branch on which we sit in among the Eukaryotes is wildly insignificant compared to all the bacterial life with which we share the planet. On the new tree, all animals and fungi – estimated to be as many as 13 million species – which fall under “opisthokonta” are found in the bottom right of the diagram, while all plant species are contained under “archaeplastida.” Knowing that really puts into perspective how the vast majority of biodiversity isn’t even visible to the human eye.

A higher resolution version is available here
The study goes further than others that have looked to redraw the tree of life by including more than 1,000 species of bacteria discovered over the last decade at UC Berkeley, which are known only by their genome. That is to say, the bacteria are so difficult to grow in the lab due to their lifestyles, which tends to involve piggybacking off other species by scavenging and parasitizing, that researchers have turned instead to DNA sequencing. They take samples from the environment, filter them, analyse all the DNA found within them, and then look for previously unknown genomes.
By doing this “metagenomic” sequencing of entire communities, the team at UC Berkeley has found hundreds of new species of bacteria in a massive range of environments, from toxic puddles in abandoned mines to the mouths of dolphins. These newly identified bacteria are often referred to as “candidate phyla radiation,” as so little is actually known about them apart from a handful of genes they contain, and can be found at the top right of the new tree. The study confirms that the candidate phyla branch forms a very major subgroup within bacteria.
“This incredible diversity means that there are a mind-boggling number of organisms that we are just beginning to explore the inner workings of that could change our understanding of biology,” says Brett Baker, another of the coauthors. This deeper understanding of the relationship between living organisms could help with other fields of science, such as in the search for new drugs.